Last updated: November 2023
Comparative genomicsQuest for orthologs
Banana split - Story
Did you know we share about 98% of our genes with chimpanzees? But how many with the banana?
Context
We share about 98% of our genes with chimpanzees. Chimpanzees are often considered as our closest ‘cousins’. That seems evident to anyone who has seen the movie Planet of the Apes: chimpanzees and humans have many characteristics in common!
But did you know that we also share some of our genes with the banana? The existence of these genes in common is proof of the relationship that exists between all living organisms.

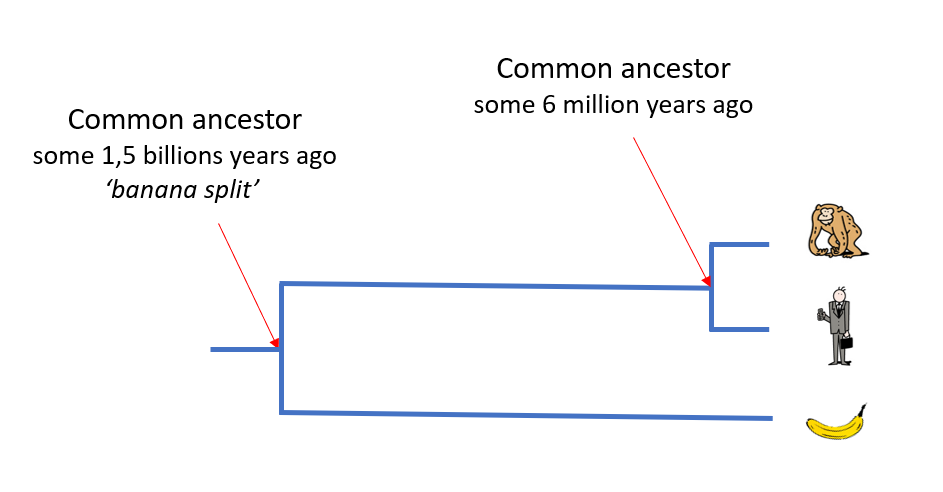
Human, chimp and banana
Humans and chimpanzees have a large proportion of genes in common, as their common ancestor lived some 6 million years ago.
The ancestors common to humans and the banana lived some 1.5 billion years ago – just before the ‘banana split’.
One can thus expect that there will be considerably fewer genes in common between humans and the banana!
What percent banana are you?
A simple search on the Internet will lead you to numerous web sites that mention that humans and the banana have between 17% and 50% of DNA in common.
Is that right? Why are the values so different? What does it mean when we refer to “percentage of DNA in common”? How can this percentage be calculated?
Source: "Neil Saunders: 50% banana" - "Do humans share 50 % of their dna with bananas?" - "The banana conjecture"

DNA in common…or genes in common?
One way would be to calculate the percentage of DNA in common between humans and banana by aligning the 2 genomes in their entirety. We could thus compare ALL the 3 billion letters A, T, C and G of the human genome sequence with ALL the 472 million letters A, T, C and G of the banana genome sequence and thus determine the percentage of similarity!
These comparisons have been made by aligning the human genome with the complete genome of other species. The results: by comparing entire genomes, the human genome is 91% similar to that of the chimpanzee, 33% similar to that of the mouse, and 1% similar to that of the zebrafish.
Source: Ensembl Compara (Human-Chimp, Human-Mouse, Human-Zebrafish)
The more distant two species are in evolution, the smaller the percentage of their genomes can be aligned. For plants, this percentage will thus be less than 1%!
Aligning the entire human genome with that of the banana (which is 6 times smaller) is thus a daunting task…and does not necessarily make sense in our case!
Why?
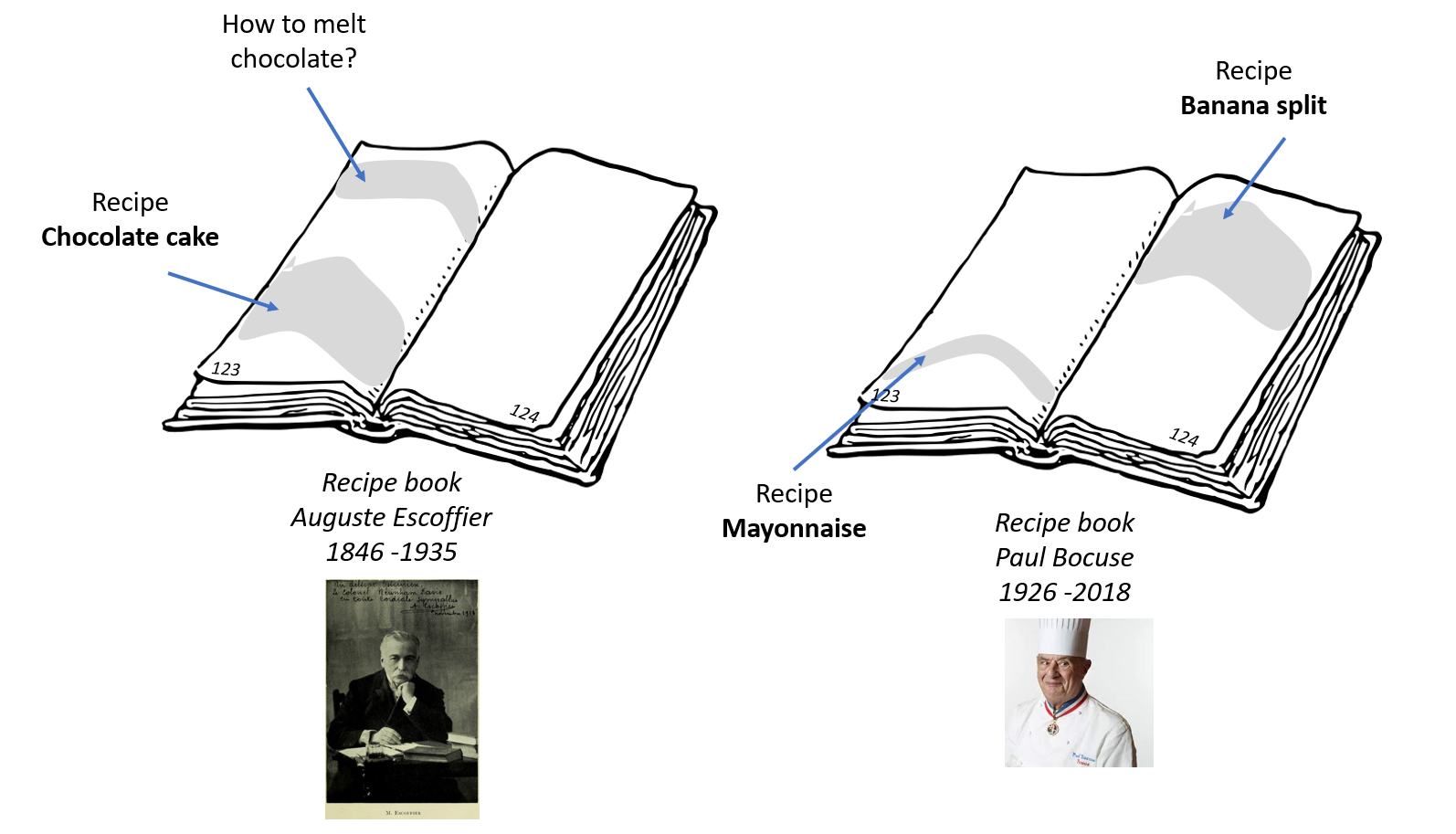
A bit like the different chapters of a recipe book, genomes are composed of different regions, which each have a specific biological function. These regions are not necessarily distributed in the same order in the genomes of different species. Some have been duplicated and some have disappeared over the course of time. And it is essential to compare what is comparable!
To come back to the recipe book, it would be useless to compare the recipe of a chocolate cake proposed by Auguste Escoffier to the recipe for mayonnaise or the Banana split of Paul Bocuse, even if they were on the same page!
But above all, all the regions of a genome do not evolve at the same rate! The regions coding for proteins, the genes – and, in particular, the regions of the genes called exons – are subject to a greater selective pressure and accumulate fewer mutations over time compared to the regions of the genome located between the genes or within introns.
The coding regions and specifically the exons are thus generally more conserved between species. They represent a very small percentage of the DNA sequence of the entire genome (less than 2% in humans), but they are comparable, and their comparison allows the highest percentage of conservation to be obtained…and this percentage of conservation is the one which makes the most sense in our story!
These calculations of the ‘% of exons DNA sequence in humans able to be aligned with the exons DNA sequence in other species (% coverage)’ have been made for the following species: humans-chimpanzee (~100 %), humans-mouse (97 %) and humans-zebrafish (58 % ).
Comparing what is comparable: an example

Here are 2 different nucleotide sequences found in the banana genome. These 2 sequences have been compared with the human genome sequence. The nucleotides that ‘map’ between human and banana are in blue.
Left: the sequence corresponds to a non-coding region of the banana genome. Mutations accumulated in non-coding regions over time have challenged the ability of bioinformatics programs to detect ‘alignable’ areas: the calculation of % similarity is very complicated.
Right: the sequence corresponds to a coding region of the banana genome, which is orthologous to the human TBB8 gene. The nucleotides in blue are correctly mapped and the calculation of the % similarity makes sense.
In order to calculate the percentage of DNA which is common to humans and the banana, we have chosen to determine the number of genes which are ‘common’ between the two species. These genes that are common between the two species are called orthologs.
The quest for orthologs between humans and banana
The programs we use for detecting orthologs do not compare the DNA sequences of genes (exons), but rather the amino acid sequences of the corresponding proteins. Each gene is associated with a ‘representative’ protein sequence which allows the number of orthologous proteins to be extrapolated to the number of orthologous genes.
We compared, two by two, 20,430 human protein sequences with 36,439 protein sequences from the banana.
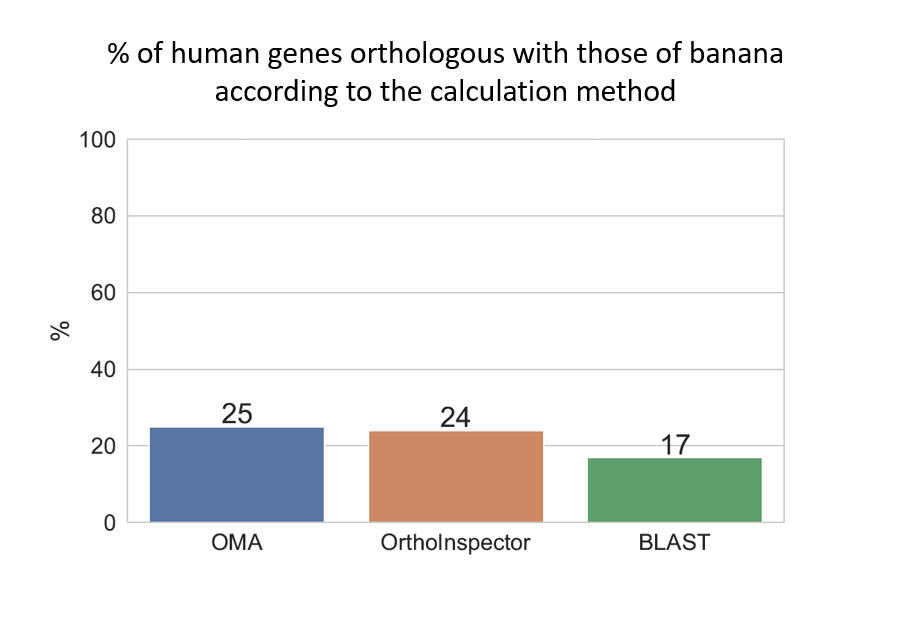
We then tested three bioinformatics methods to find orthologs: OMA, OrthoInspector, and BLAST.
In all of the bioinformatics methods used, the similarity between the sequences (and several statistical criteria) is what determines if two genes in different species are orthologs.
Here is a part of alignment between a human protein and a protein of the banana which are ‘orthologs’ (gene TUBB8). Find the differences! (source)
We then divided the number of orthologs found by the total number of genes in the genome of each of the 2 species and took the average to obtain the percentage of shared genes.
An example of calculation:
Each circle represents ~ 1,000 genes.
The yellow circles represent the genes shared between humans and the banana.
Question:
1. What % of human genes are common to genes in the banana?
2. What % of banana genes are common to genes in humans?
Any one gene can be present in multiple copies in another species: that’s the reason why the number of yellow circles (orthologs) is not the same in humans and in the banana!
Answer
5,000 genes / 20,000, that is 25 % of human genes are ‘common’ to those in the banana.
9,000 genes / 36,000, that is 25 % of banana genes are ‘common’ to those in humans.
For the calculations, we have chosen to take the average of these 2 percentages.
Results obtained by the experts
Depending on the method used, between 3,400 et 4,900 genes are common to both humans and the banana. That represents between 17% and 25% of human genes.
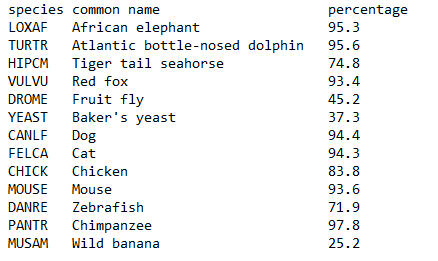
We have 98% of genes in common with the chimpanzee, 94% of genes in common with the mouse, 72% of genes in common with the zebrafish …. And thus about 25% of genes in common with the banana (OMA method).
Although this percentage of conservation has been calculated analyzing less than 2% of the human genome (coding regions/exons), this number is not negligible!
That means that the human/banana orthologous genes have been conserved for 1.5 billion years of evolution!
What is the function of human/banana orthologs?
We looked at the function associated with each of the orthologous genes. This information is found in a specialised database, such as UniProtKB.
Unsurprisingly, the proteins that humans and the banana have in common are primarily implicated in basic metabolic processes, such as gene expression, lipid metabolism, or the modification of RNA (splicing).
Even though humans and bananas 'split' 1.5 billion years ago, remarkably 25 % of our genes remain similar. These shared genes are responsible for the biological functions essential to the life of all eukaryotic organisms. That's bananas!